RESEARCH
Understanding how cells grow and divide has profound impacts on basic science, biotechnology, and medicine. Despite recent advances in molecular biology and biochemistry, a central challenge remains: bridging the nanometer-scale activities of proteins and the construction of entire cells. Although the mechanisms of bacterial proliferation have been a major focus of research for over a century, it has remained difficult to determine how cellular structure and organization are dynamically controlled due to the central—yet neglected—importance of physical factors.
To address these knowledge gaps, our lab pursues research directions that span from the atomic to the multicellular scales. We investigate the physical nature of intracellular spatial organization, mechanics, and kinetics by leveraging top-down approaches based on cellular-scale observations, bottom-up approaches based on biophysical molecular observations, and computational modeling that connects the two paradigms. Understanding cellular growth and form remains a fascinating, multifaceted challenge with obvious implications for health and disease. In addition to the importance of bacteria as a model system for basic science, uncovering the general physical rules that underlie how bacteria grow and divide will have important applications for controlling bacterial communities and developing novel strategies in synthetic biology.
| Research Topics |
|---|
| Systems biology of essential genes in growth |
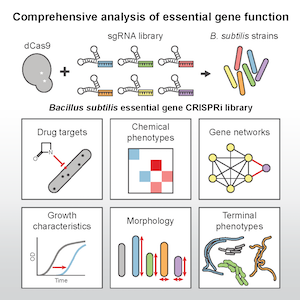 Essential genes underpin core cellular processes required for survival. However, little is known about the genetic interactions among essential processes and the physiological effects of depleting these processes. Elucidating these relationships is critical for understanding and manipulating bacterial growth. This knowledge is also crucial for drug development, as most drugs target essential processes. To comprehensively assess essential gene function, we are collaborating with the Gross Lab at UCSF and utilizing a library of CRISPRi-mediated knockdowns of every essential gene in various organisms to determine phenotypes during chemical treatment, population growth, and single-cell behavior during essential gene depletion. Using high-throughput microscopy and chemical genomics, we have uncovered a network rich in novel connections among essential processes required for cell viability. We have also shown that the “essentiality” of a gene is highly dependent on the phase of growth and cell morphology. Depletion of essential genes provides single-cell death phenotypes that correlate strongly with gene function and antibiotic targeting. Moreover, we have demonstrated that CRISPRi can be used to dissect interactions between among multiple proteins, such as a triplet of penicillin binding proteins whose joint interaction is essential for effective cell division and elongation. This strategy for analysis of essential genes is generally applicable to diverse bacteria and should empower the greater biomedical and fundamental research communities.
Essential genes underpin core cellular processes required for survival. However, little is known about the genetic interactions among essential processes and the physiological effects of depleting these processes. Elucidating these relationships is critical for understanding and manipulating bacterial growth. This knowledge is also crucial for drug development, as most drugs target essential processes. To comprehensively assess essential gene function, we are collaborating with the Gross Lab at UCSF and utilizing a library of CRISPRi-mediated knockdowns of every essential gene in various organisms to determine phenotypes during chemical treatment, population growth, and single-cell behavior during essential gene depletion. Using high-throughput microscopy and chemical genomics, we have uncovered a network rich in novel connections among essential processes required for cell viability. We have also shown that the “essentiality” of a gene is highly dependent on the phase of growth and cell morphology. Depletion of essential genes provides single-cell death phenotypes that correlate strongly with gene function and antibiotic targeting. Moreover, we have demonstrated that CRISPRi can be used to dissect interactions between among multiple proteins, such as a triplet of penicillin binding proteins whose joint interaction is essential for effective cell division and elongation. This strategy for analysis of essential genes is generally applicable to diverse bacteria and should empower the greater biomedical and fundamental research communities.
A Comprehensive, CRISPR-based Functional Analysis of Essential Genes in Bacteria.
Peters JM*, Colavin A*, Shi H*, Czarny TL, Larson MH, Wong S, Hawkins JS , Lu CHS, Koo B-M , Marta E, Shiver AL, Whitehead EH, Weissman JS, Brown ED, Qi LS†, Huang KC†, Gross CA†. Cell. 2016 165 1-14 (2016).
*Co-first authors †Co-corresponding authors