RESEARCH
Understanding how cells grow and divide has profound impacts on basic science, biotechnology, and medicine. Despite recent advances in molecular biology and biochemistry, a central challenge remains: bridging the nanometer-scale activities of proteins and the construction of entire cells. Although the mechanisms of bacterial proliferation have been a major focus of research for over a century, it has remained difficult to determine how cellular structure and organization are dynamically controlled due to the central—yet neglected—importance of physical factors.
To address these knowledge gaps, our lab pursues research directions that span from the atomic to the multicellular scales. We investigate the physical nature of intracellular spatial organization, mechanics, and kinetics by leveraging top-down approaches based on cellular-scale observations, bottom-up approaches based on biophysical molecular observations, and computational modeling that connects the two paradigms. Understanding cellular growth and form remains a fascinating, multifaceted challenge with obvious implications for health and disease. In addition to the importance of bacteria as a model system for basic science, uncovering the general physical rules that underlie how bacteria grow and divide will have important applications for controlling bacterial communities and developing novel strategies in synthetic biology.
| Research Topics |
|---|
| The structural dynamics of cytoskeletal proteins |
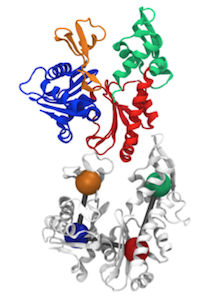 How does a protein such as MreB detect micron-scale geometry, exert forces on the membrane (Wang et al., 2010), and dictate cellular chirality (Wang et al., 2012)? Since every atom matters at the molecular scale, we used all-atom molecular dynamics simulations to determine that ATP-bound MreB monomers rapidly adopt a torqued configuration in a polymerization-dependent manner, similar to actin (Colavin et al., 2014) that controls filament bending. Our simulations yielded an unprecedented understanding of cellular morphogenesis, directly connecting geometric sensing by MreB filaments (Ursell et al., 2014) to supmolecular transitions responsible for curvature and flattening. This understanding is empowering an atomic-scale framework for predicting cellular phenotypes, which is particularly important for the rational design of molecular inhibitors.
How does a protein such as MreB detect micron-scale geometry, exert forces on the membrane (Wang et al., 2010), and dictate cellular chirality (Wang et al., 2012)? Since every atom matters at the molecular scale, we used all-atom molecular dynamics simulations to determine that ATP-bound MreB monomers rapidly adopt a torqued configuration in a polymerization-dependent manner, similar to actin (Colavin et al., 2014) that controls filament bending. Our simulations yielded an unprecedented understanding of cellular morphogenesis, directly connecting geometric sensing by MreB filaments (Ursell et al., 2014) to supmolecular transitions responsible for curvature and flattening. This understanding is empowering an atomic-scale framework for predicting cellular phenotypes, which is particularly important for the rational design of molecular inhibitors.
Structural basis for the geometry-driven localization of a small protein.
Gill RL Jr*, Castaing JP*, Hsin J*, Tan IS, Wang X, Huang KC†, Tian F†, Ramamurthi KS†.
Proc Natl Acad Sci U S A. 2015 Apr 14;112(15):E1908-15. doi: 10.1073/pnas.1423868112. Epub 2015 Mar 30.
*Co-first authors †Co-corresponding authors
Variations in the binding pocket of an inhibitor of the bacterial division protein FtsZ across genotypes and species.
Miguel A, Hsin J, Liu T, Tang G, Altman RB, Huang KC.
PLoS Comput Biol. 2015 Mar;11(3):e1004117. doi: 10.1371/journal.pcbi.1004117.
Effects of polymerization and nucleotide identity on the conformational dynamics of the bacterial actin homolog MreB.
Colavin A, Hsin J, Huang KC.
Proc Natl Acad Sci U S A. 2014 Mar 4;111(9):3585-90. doi: 10.1073/pnas.1317061111. Epub 2014 Feb 18.
FtsZ protofilaments use a hinge-opening mechanism for constrictive force generation.
Li Y, Hsin J*, Zhao L*, Cheng Y*, Shang W, Huang KC, Wang HW, Ye S.
Science. 2013 Jul 26;341(6144):392-5. doi: 10.1126/science.1239248.
*Co-first authors
Posttranslational acetylation of α-tubulin constrains protofilament number in native microtubules.
Cueva JG, Hsin J, Huang KC, Goodman MB.
Curr Biol. 2012 Jun 19;22(12):1066-74. doi: 10.1016/j.cub.2012.05.012. Epub 2012 May 31.
Nucleotide-dependent conformations of FtsZ dimers and force generation observed through molecular dynamics simulations
Hsin J, Gopinathan A, Huang KC. Proc Natl Acad Sci U S A. 2012 Jun 12;109(24):9432-7. doi: 10.1073/pnas.1120761109. Epub 2012 May 30.