RESEARCH
Understanding how cells grow and divide has profound impacts on basic science, biotechnology, and medicine. Despite recent advances in molecular biology and biochemistry, a central challenge remains: bridging the nanometer-scale activities of proteins and the construction of entire cells. Although the mechanisms of bacterial proliferation have been a major focus of research for over a century, it has remained difficult to determine how cellular structure and organization are dynamically controlled due to the central—yet neglected—importance of physical factors.
To address these knowledge gaps, our lab pursues research directions that span from the atomic to the multicellular scales. We investigate the physical nature of intracellular spatial organization, mechanics, and kinetics by leveraging top-down approaches based on cellular-scale observations, bottom-up approaches based on biophysical molecular observations, and computational modeling that connects the two paradigms. Understanding cellular growth and form remains a fascinating, multifaceted challenge with obvious implications for health and disease. In addition to the importance of bacteria as a model system for basic science, uncovering the general physical rules that underlie how bacteria grow and divide will have important applications for controlling bacterial communities and developing novel strategies in synthetic biology.
| Research Topics |
|---|
| Modeling growth of the entire bacterial cell wall |
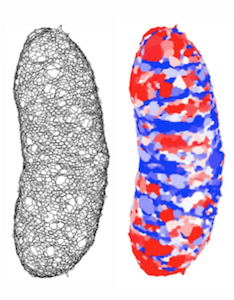 Bacteria come in a wide variety of shapes and sizes that play a critical role in important functions such as attachment, dispersal, motility, and pathogenesis. The cell wall is both necessary and sufficient for determining shape, as it bears the stress generated by large turgor pressures. Thus, bacterial morphogenesis is a self-evident physical phenomenon, ultimately involving a mechanochemical blueprint that balances the spatiotemporal program of growth with physical forces of expansion. The wide disparity between the length and time scales of the molecular machinery and those of growth requires a computational model that encompasses spatial patterning, biochemical regulation, and physical perturbations over time scales of seconds to hours. Our group developed the first quantitative, 3D physical model of the cell wall that predicts the response of cell shape to wall damage and other perturbations (Huang et al., 2008). We successfully applied this model to reveal the consequence of antibiotics on cell shape (Huang et al., 2008), principles for robust maintenance of shape (Furchtgott et al., 2011), the intrinsic dynamics of the cytoskeleton during cell-wall synthesis (Teeffelen et al., 2011), and the establishment of chirality in cell-wall organization (Wang et al., 2012). Each time, our success generated new strategies for intervening in cell growth, with major therapeutic possibilities.
Bacteria come in a wide variety of shapes and sizes that play a critical role in important functions such as attachment, dispersal, motility, and pathogenesis. The cell wall is both necessary and sufficient for determining shape, as it bears the stress generated by large turgor pressures. Thus, bacterial morphogenesis is a self-evident physical phenomenon, ultimately involving a mechanochemical blueprint that balances the spatiotemporal program of growth with physical forces of expansion. The wide disparity between the length and time scales of the molecular machinery and those of growth requires a computational model that encompasses spatial patterning, biochemical regulation, and physical perturbations over time scales of seconds to hours. Our group developed the first quantitative, 3D physical model of the cell wall that predicts the response of cell shape to wall damage and other perturbations (Huang et al., 2008). We successfully applied this model to reveal the consequence of antibiotics on cell shape (Huang et al., 2008), principles for robust maintenance of shape (Furchtgott et al., 2011), the intrinsic dynamics of the cytoskeleton during cell-wall synthesis (Teeffelen et al., 2011), and the establishment of chirality in cell-wall organization (Wang et al., 2012). Each time, our success generated new strategies for intervening in cell growth, with major therapeutic possibilities.
Helical insertion of peptidoglycan produces chiral ordering of the bacterial cell wall.
Wang S, Furchtgott L, Huang KC†, Shaevitz JW†.
Proc Natl Acad Sci U S A. 2012 Mar 6;109(10):E595-604. doi: 10.1073/pnas.1117132109. Epub 2012 Feb 17.
†Co-corresponding authors
The bacterial actin MreB rotates, and rotation depends on cell-wall assembly.
van Teeffelen S, Wang S, Furchtgott L, Huang KC, Wingreen NS, Shaevitz JW, Gitai Z.
Proc Natl Acad Sci U S A. 2011 Sep 20;108(38):15822-7. doi: 10.1073/pnas.1108999108. Epub 2011 Sep 8.
Mechanisms for maintaining cell shape in rod-shaped Gram-negative bacteria.
Furchtgott L, Wingreen NS, Huang KC.
Mol Microbiol. 2011 Jul;81(2):340-53. doi: 10.1111/j.1365-2958.2011.07616.x. Epub 2011 Apr 18.
Cell shape and cell-wall organization in Gram-negative bacteria.
Huang KC, Mukhopadhyay R, Wen B, Gitai Z, Wingreen NS.
Proc Natl Acad Sci U S A. 2008 Dec 9;105(49):19282-7. doi: 10.1073/pnas.0805309105. Epub 2008 Dec 2.